Gallocatechin from Uncaria gambir Roxb as a MurB Inhibitor: A Molecular Docking Analysis and Its Therapeutic Implications
Research Article
DOI:
https://doi.org/10.33005/jdiversemedres.v3i3.322Keywords:
Antibacterial agents, flavonoids, gallocatechin, molecular docking, MurB enzyme, peptidoglycan biosynthesis, Pseudomonas aeruginosaAbstract
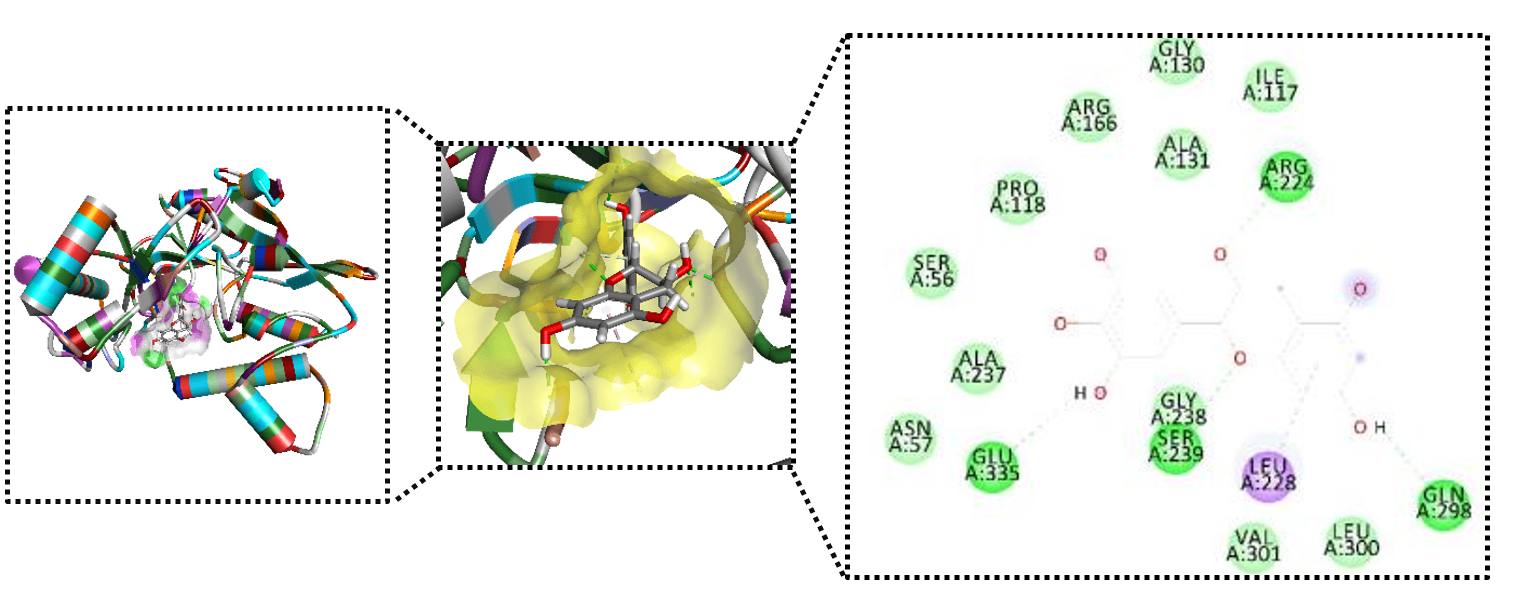
The rapid escalation of antimicrobial resistance demands new antibacterial strategies targeting essential and druggable bacterial enzymes. Here, we report the molecular characterization of gallocatechin, a polyphenolic compound from Uncaria gambir, as a potential inhibitor of MurB from Pseudomonas aeruginosa (PDB ID: 7ORZ), a key enzyme in peptidoglycan biosynthesis. Docking protocol validation via redocking of the co-crystallized ligand yielded high structural accuracy (RMSD = 0.991 Å). Gallocatechin exhibited a markedly enhanced binding affinity (−8.1 kcal/mol) relative to the reference ligand (−5.3 kcal/mol), corresponding to an approximately 100-fold lower predicted inhibition constant (Ki ≈ 1.09 µM vs 131 µM). Structural analysis revealed that gallocatechin establishes a dense and multi-modal interaction network, simultaneously engaging the catalytic triad (Arg166, Ser239, Glu335) through hydrogen bonding and complementary electrostatic interactions. Notably, this tri-residue engagement and dual electrostatic stabilization are rarely observed in previously reported MurB inhibitors. The binding mode supports a dual inhibitory mechanism involving both competitive substrate displacement and perturbation of the NADPH-dependent catalytic cycle. Collectively, these findings position gallocatechin as a structurally distinct and mechanistically promising scaffold for MurB inhibition, providing a rational basis for the development of next-generation antibacterial agents targeting multidrug-resistant P. aeruginosa. Further experimental validation is warranted to confirm its therapeutic potential.
Downloads
References
Santos C, Rodrigues GR, Lima LF, dos Reis MCG, Cunha NB, Dias SC, et al. Advances and perspectives for antimicrobial peptide and combinatory therapies. Front Bioeng Biotechnol. 2022;10.
Souza PFN, Filho NS dos S, Mororó JLT, Brito DM da S, da Lima AB, Mesquita FP, et al. Pandemic Events Caused by Bacteria Throughout Human History and the Risks of Antimicrobial Resistance Today. Microorganisms. 2025;13(2):457.
Piscitelli R, Iula DV, Birra D, Pandolfi V, Nuzzo V, Papa M, et al. Antimicrobial Stewardship: When Less Is More [Internet]. 2024. Available from: https://www.preprints.org/manuscript/202407.1386/v1
Ablakimova N, Smagulova GA, Rachina S, Mussina AZ, Zare A, Mussin NM, et al. Bibliometric Analysis of Global Research Output on Antimicrobial Resistance among Pneumonia Pathogens (2013–2023). Antibiotics. 2023;12(9):1411.
Renz J, Dauda KA, Aga ONL, Diaz-Uriarte R, Löhr IH, Blomberg B, et al. Evolutionary accumulation modeling in AMR: machine learning to infer and predict evolutionary dynamics of multidrug resistance. MBio. 2025;16(6).
Prasetyoputri A. Detection of Bacterial Coinfection in COVID-19 Patients Is a Missing Piece of the Puzzle in the COVID-19 Management in Indonesia. ACS Infect Dis. 2021;7(2):203–5.
Krockow E, Jones M, Mkumbuzi S, Mendelson M, Tarrant C, Froud R, et al. Developing public health risk messages about antibiotic resistance using metaphors: An international co-design and e-Delphi consensus study [Internet]. 2025. p. 1–31. Available from: https://www.researchsquare.com/article/rs-7602040/v1
Fukuda D, Handa Y, Kayama Y, Fujii K, Kawamatsu S, Kawano Y, et al. The Current Landscape of Antibiotic Use and Antimicrobial Resistance in Japan: Focusing on Common Infections Including Uncomplicated Urinary Tract Infection and Gonorrhea. Antibiotics. 2025;14(8):813.
Jeong Y Il, Lee HY, Lee S, Jeong GY, Kim SH, Kim S, et al. Korea’s National Action Plan on Antimicrobial Resistance: Focusing on the Appropriate Use of Antibiotics. Infect Chemother. 2025;57(2):203–14.
Mir TA, Shareef T, Lone SA, Mir SA, Ahmad J, Ganai BA. Antibiotic Resistance Profiling and Identification of Risk Factors Associated With Prevalence of Urinary Tract Infections: A Cross-Sectional Study. Apmis. 2025;133(10).
Silvestro S, Biondo C, Midiri A, Lucia B, Mancuso G. The Role of Livestock Antibiotic Use in Microbiota Dysbiosis and Neuroinflammation. Antibiotics. 2025;14(6):608.
Tuytschaevers S, Aden L, Greene Z, Nixon C, Shaw W, Hatch D, et al. Isolation, whole-genome sequencing, and annotation of two antibiotic-producing and antibiotic-resistant bacteria, Pantoea rodasii RIT 836 and Pseudomonas endophytica RIT 838, collected from the environment. PLoS One. 2024;19(2 FEBRUARY):e0293943.
Kurniawan I, Rohmatika AU. Molecular Docking, QSAR, and Bioactivity Prediction of Uncaria gambir Flavonoids as Antibacterial Agents Targeting MurA Enzyme. Biointerface Res Appl Chem. 2026;16(1):1–22.
Ilham Kurniawan. Biokimia pertanian: dasar molekuler untuk produktivitas dan keberlanjutan (Edisi 1). 1st ed. Wanti Mindari, editor. Lamongan: Ghani Press; 2026. 110 p.
Kurniawan I, Winarno NS. Unlocking Antibacterial Potential: Thiophene-2-carbaldehyde Modification of Acertannin from African Leaves as MurA Enzyme Inhibitors. J Ners. 2025;9(4):7602–12.
Kurniawan I, Ambarsari L, Kurniatin P, Tri Wahyudi S. Novel Compounds Design of Acertannin, Hamamelitannin, and Petunidin-3-Glucoside Typical Compounds of African Leaves (Vernonia amygdalina Del) as Antibacterial Based on QSAR and Molecular Docking. J Jamu Indones. 2024;8:29–38.
Kurniawan I. Modifikasi Struktur Senyawa Tanin Daun Afrika sebagai Antibakteri pada Target MurA dengan Metode QSAR dan Komputasi Dinamika Molekul. Bogor (ID): IPB University; 2022.
Kumar V, Shetty P, Arunodaya HS, Chandra K. S, Ramu R, Patil SM, et al. Potential Fluorinated Anti-MRSA Thiazolidinone Derivatives with Antibacterial, Antitubercular Activity and Molecular Docking Studies. Chem Biodivers. 2022;19(2).
Kurniawan I. Analisis Penambatan Molekuler Turunan Senyawa Tanin Daun Afrika (Vernonia amygdalina Del) terhadap MurA sebagai Antibakteri. Bogor (ID); IPB University; 2021.
Acebrón-Garciá-De-Eulate M, Mayol-Llinàs J, Holland MTO, Kim SY, Brown KP, Marchetti C, et al. Discovery of Novel Inhibitors of Uridine Diphosphate- N-Acetylenolpyruvylglucosamine Reductase (MurB) from Pseudomonas aeruginosa, an Opportunistic Infectious Agent Causing Death in Cystic Fibrosis Patients. J Med Chem. 2022;65(3):2149–73.
Haque MA, Singh M, Tripathi MK, Ethayathulla AS, Kaur P. Identification of natural small molecule modulators of MurB from Salmonella enterica serovar Typhi Ty2 strain using computational and biophysical approaches. Proteins Struct Funct Bioinforma. 2023;91(3):363–79.
Tratrat C, Petrou A, Geronikaki A, Ivanov M, Kostić M, Soković M, et al. Thiazolidin-4-Ones as Potential Antimicrobial Agents: Experimental and In Silico Evaluation. Molecules. 2022;27(6):1930.
Isa MA, Mohammed MM. Molecular docking and dynamic simulation of UDP-N-acetylenolpyruvoylglucosamine reductase (MurB) obtained from Mycobacterium tuberculosis using in silico approach. Netw Model Anal Heal Informatics Bioinforma. 2021;10(1).
Masumi M, Noormohammadi F, Kianisaba F, Nouri F, Taheri M, Taherkhani A. Methicillin-Resistant Staphylococcus aureus: Docking-Based Virtual Screening and Molecular Dynamics Simulations to Identify Potential Penicillin-Binding Protein 2a Inhibitors from Natural Flavonoids. Int J Microbiol. 2022;2022:1–14.
Mehta K, Khambete M, Abhyankar A, Omri A. Anti-Tuberculosis Mur Inhibitors: Structural Insights and the Way Ahead for Development of Novel Agents. Pharmaceuticals. 2023;16(3):377.
Verma A, Kumar V, Naik B, Masood Khan J, Singh P, Erik Joakim Saris P, et al. Screening and molecular dynamics simulation of compounds inhibiting MurB enzyme of drug-resistant Mycobacterium tuberculosis: An in-silico approach. Saudi J Biol Sci. 2023;30(8):103730.
Kurniawan I, Ambarsari L, Kurniatin PA, Wahyudi ST. Novel Compounds Design of Acertannin, Hamamelitannin, and Petunidin-3-Glucoside Typical Compounds of African Leaves (Vernonia amygdalina Del) as Antibacterial Based on QSAR and Molecular Docking. J Jamu Indones. 2024;8(2):29–38.
Kashyap P, Verma S, Gupta P, Narang R, Lal S, Devgun M. Recent insights into antibacterial potential of benzothiazole derivatives. Med Chem Res. 2023;32(8):1543–73.
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Journal of Diverse Medical Research : Medicosphere

This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
CC Attribution 4.0

